Transcriptional Target Selection
Curating lists of genes to target with Transcription Factors in Neurodegeneration
# imports
import os, sys
import io
import json
import requests
def flatten(lol): return [x for l in lol for x in l]
def chunker(seq, size): return (seq[pos:pos + size] for pos in range(0, len(seq), size))
import pandas as pd
import numpy as np
import networkx as nx
import matplotlib.pyplot as plt
import seaborn as sns#; sns.set_theme()
plt.rcParams['figure.figsize'] = [12, 5]
plt.rcParams['figure.dpi'] = 140
plt.rcParams['agg.path.chunksize'] = 10000
%config InlineBackend.figure_format = 'retina'
%matplotlib inline
import matplotlib_inline
matplotlib_inline.backend_inline.set_matplotlib_formats('svg')
from IPython.display import Image, HTML, IFrame, SVG, display
from Bio import SeqIO
from Bio.KEGG import REST, Gene
from Bio.KEGG.KGML import KGML_parser
from Bio.Graphics.KGML_vis import KGMLCanvas
from wand.image import Image as WImage
def wImage(fn, **kwargs): return WImage(filename=fn, **kwargs)
pd.set_option('display.max_rows', 200)
# venn diagrams
import matplotlib.pyplot as plt
from matplotlib_venn import venn2, venn3
import upsetplot
import matplotlib_inline.backend_inline
matplotlib_inline.backend_inline.set_matplotlib_formats('svg')
%config InlineBackend.figure_format = 'retina'
%matplotlib inline
plt.rcParams['figure.figsize'] = [6, 3]
plt.rcParams['figure.dpi'] = 140
plt.rcParams['font.size'] = 8
import re
import traceback
def remove_special_characters(s): return re.sub('[^a-zA-Z0-9]', ' ', s)
def get_strings(stack):
filename, lineno, function_name, code = stack[-2]
names = [n.strip() for n in code[code.find("(")+1:code.rfind(")")].split(',')]
common_prefix = os.path.commonprefix(names)
common_suffix = os.path.commonprefix([name[::-1] for name in names])[::-1]
if len(common_prefix) > 2:
names = [name.removeprefix(common_prefix) for name in names]
if len(common_suffix) > 2:
names = [name.removesuffix(common_suffix) for name in names]
if len(common_prefix) > 2 or len(common_suffix) > 2:
names = [remove_special_characters(name) for name in names]
return names
def form_list_of_tuples(names, iterables):
final = []
for name, iterable in zip(names, iterables):
len_full = len(iterable)
s = set(iterable)
len_uniques = len(s)
if len_uniques == len_full:
name += f' ({len_uniques})'
else:
name += f' ({len_uniques} unique in {len_full})'
final.append((s, name))
return final
def venn(*args, **kwargs):
if len(args) == 2: method = venn2
elif len(args) == 3: method = venn3
else: print('incorrect number of args')
names = get_strings(traceback.extract_stack())
final = form_list_of_tuples(names, args)
return method(*zip(*final), **kwargs)
def upset(*args, **kwargs):
names = get_strings(traceback.extract_stack())
final = form_list_of_tuples(names, args)
s = upsetplot.from_contents(dict(final))
fig, upsetplot.UpSet(s, subset_size='count', show_counts=True).plot()
return s, fig
# clustering heatmaps
import scipy
import scipy.stats
from scipy.cluster.hierarchy import dendrogram, linkage
from scipy.cluster import hierarchy
def sort_df_by_hclust_olo(df, how='both', method='ward', metric='euclidean'):
'''
how={'index', 'columns', 'both'}
'''
df = df.fillna(0)
if how in ['index', 'both']:
Z = linkage(df, method=method, metric=metric)
order = hierarchy.leaves_list(hierarchy.optimal_leaf_ordering(Z, df))
df = df.iloc[order]
if how in ['columns', 'both']:
df = df.T
Z = linkage(df, method=method, metric=metric)
order = hierarchy.leaves_list(hierarchy.optimal_leaf_ordering(Z, df))
df = df.iloc[order].T
return df.replace(0, np.nan)
# scaling SVG
from IPython.display import SVG
from bs4 import BeautifulSoup
import re
def scale_svg(svg_object, scale=1.0):
soup = BeautifulSoup(svg_object.data, 'lxml')
svg_elt = soup.find("svg")
w = svg_elt.attrs["width"].rstrip("pt")
h = svg_elt.attrs["height"].rstrip("pt")
ws = float(w)*scale
hs = float(h)*scale
svg_elt.attrs["width"] = f"{ws}pt"
svg_elt.attrs["height"] = f"{hs}pt"
svg_elt.attrs["viewbox"] = f"0.00 0.00 {ws} {hs}"
g_elt = svg_elt.find("g")
tf = g_elt.attrs["transform"]
# non-greedy regex-search-and-replace
tf2 = re.sub(
"scale\(.*?\)",
f"scale({scale} {scale})",
tf
)
g_elt.attrs["transform"] = tf2
svg_object.data = str(svg_elt)
return svg_object
TODO in the future:
KEGG functions
# load reference genome and KEGG IDs
id_mapping = pd.read_csv('/Users/alex/Documents/scImputation/KEGG_hsa_ncbi_and_ENSG_ID_map.csv')
genome = pd.read_csv('/Users/alex/Documents/AChroMap/data/processed/GRCh38_V40.csv')
genome['gene_id'] = genome.gene_id.str.split('.').str[0]
gene_names = genome[['gene_id', 'gene_name']].drop_duplicates('gene_id').set_index('gene_id')
# ko_to_hsa = pd.read_csv('https://rest.kegg.jp/link/ko/hsa', sep='\t', header=None, names=['hsa', 'ko'])
# ko_to_hsa.to_csv('ko_to_hsa.csv', index=False)
# id_mapping = id_mapping.merge(ko_to_hsa, how='left', left_on='KEGG_id', right_on='hsa')
# define functions to query KEGG database and process/visualize results
def to_df(result, **kwargs):
return pd.read_table(io.StringIO(result), header=None, **kwargs)
def kegg_find(query):
return to_df(REST.kegg_find('pathway', query).read(), names=['pathway_ID', 'pathway_name'])
def kegg2pdf(map_id, fn=None):
# Get the background image first
pathway = KGML_parser.read(REST.kegg_get(map_id, "kgml"))
canvas = KGMLCanvas(pathway, import_imagemap=True)
if isinstance(fn, type(None)):
img_filename = "%s.pdf" % map_id
else:
img_filename = fn
canvas.draw(img_filename)
return img_filename
def get_KEGG_pathway_gene_members(pathway):
pathway = KGML_parser.read(REST.kegg_get(pathway, 'kgml'))
members = np.unique(flatten([gene.name.strip().split(' ') for gene in pathway.genes]))
members = id_mapping[id_mapping.KEGG_id.isin(members)].drop_duplicates('KEGG_id').set_index('KEGG_id').reindex(members)
members['gene_name'] = gene_names.reindex(members.ENSG).gene_name.values
members = members[['ENSG', 'gene_name']].dropna()
return members
def unpack_BRITE_json(d):
if 'children' in d:
return {d['name']: [unpack_BRITE_json(child) for child in d['children']]}
else:
if d['name'].startswith('K'):
tmp = d['name'].split(';')
tmp0 = tmp[0].split(' ')
return [tmp0[0], [s.strip() for s in tmp0[1].split(',')], tmp[1].strip()]
else:
return d['name']
def second_parse(d, index=[]):
if isinstance(d, dict):
return flatten([second_parse(x, index+[k]) for k,v in d.items() for x in v])
else:
# print(d)
return [[index, d[0], d[1]]]
all_gene_symbols_set = set(gene_names.gene_name.values)
def get_KEGG_BRITE_entry(brite_id):
r = requests.get(f'https://rest.kegg.jp/get/br:{brite_id}/json')
j = unpack_BRITE_json(r.json())
j2 = pd.DataFrame(second_parse(j))
j2 = j2.merge(id_mapping, how='left', left_on=1, right_on=id_mapping.ko.str.split(':').str[1])
j2.index = pd.MultiIndex.from_tuples([tuple(x) for x in j2[0].values])
brite_id = brite_id.split('+')[0]
j2 = j2.loc[brite_id].drop(['ncbi_ID','KEGG_id','ko'], axis=1).rename(columns={1:'KEGG_ortholog_ID',2:'KEGG_gene_name'})
j2['KEGG_gene_name'] = j2.KEGG_gene_name.apply(lambda l: [x for x in l if x in all_gene_symbols_set])
j2['KEGG_gene_name'] = j2.KEGG_gene_name.apply(lambda l: l if len(l) else np.nan)
j2['gene_name'] = gene_names.reindex(j2.ENSG).values
j2 = j2.dropna(subset=['KEGG_gene_name','ENSG','gene_name'], how='all')
def newcol(row):
if (isinstance(row.KEGG_gene_name, list) and row.gene_name in row.KEGG_gene_name) or (str(row.KEGG_gene_name) == 'nan'):
return np.nan
else:
return row.KEGG_gene_name[0]
j2['newcol'] = j2.apply(newcol, axis=1)
save_index = j2.index
j2 = j2.merge(gene_names.reset_index(), how='left', left_on='newcol', right_on='gene_name')
j2.index = save_index
j2['gene_name_x'] = j2.gene_name_x.fillna(j2.gene_name_y)
j2['ENSG'] = j2.ENSG.fillna(j2.gene_id)
j2 = j2.drop(['KEGG_gene_name', 'newcol', 'gene_id', 'gene_name_y', 0, 'KEGG_ortholog_ID'], axis=1).rename(columns={'gene_name_x': 'gene_name'})
return j2
def get_KEGG_data(pathway_code=None, brite_code=None):
pathway_genes = None
brite_genes = None
if pathway_code:
Image(REST.kegg_get(pathway_code, 'image').read())
pathway_genes = get_KEGG_pathway_gene_members(pathway_code)
if brite_code:
brite_genes = get_KEGG_BRITE_entry(brite_code)
if pathway_code:
brite_genes['in_KEGG_pathway'] = brite_genes.ENSG.isin(pathway_genes.ENSG).astype(int)
return pathway_genes, brite_genes
GO functions
# define functions to query GO database and process/visualize results
def GO_subgraph_from_query(query):
# nodes = pd.DataFrame({key: data for key, data in ontology_graph.nodes(data=True) if all(query in data['name'].lower() for query in queries)}, index=GO_columns).T
nodes = pd.DataFrame({key: data for key, data in ontology_graph.nodes(data=True) if query.lower() in data['name'].lower()}, index=GO_columns).T
nodes['gene_name'] = nodes.ENSG.apply(lambda l: gene_names.reindex(l).gene_name.values)
g = ontology_graph_literate.subgraph(nodes.name.tolist())
return g, nodes
def draw_graph(graph, scale=0.5):
ag = nx.nx_agraph.to_agraph(graph)
ag.graph_attr['rankdir']='LR'
ag.node_attr['shape'] = 'record'
svg = ag.draw(prog='dot',format='svg')
# display(scale_svg(SVG(svg), scale))
display(SVG(svg))
def create_GO_by_gene_matrix(node_df, attr='ENSG'):
df = node_df.set_index('name')[attr].explode()
df = (df.reset_index()
.assign(v = True)
.groupby(['name',attr]).first()
.unstack(level=1)
.fillna(False)
)['v']
if attr == 'ENSG': df = df.loc[:, df.columns.str.startswith('gene_name')]
return df
sns.set(font_scale = 0.5)
cmap = sns.color_palette("light:b", as_cmap=True)
def plot_gene_membership_in_GO_terms(df, figsize=(10, 50)):
ax = sns.clustermap(df.T, figsize=figsize, cbar=False, dendrogram_ratio=(0.001,0.001), colors_ratio=0.0001, cmap=cmap)
ax.ax_row_dendrogram.set_visible(False)
ax.ax_col_dendrogram.set_visible(False)
ax.ax_cbar.set_visible(False)
plt.xticks(fontsize=8)
plt.yticks(fontsize=8)
return ax
ontology_graph = nx.read_gpickle('/Users/alex/Documents/goenrich-web/GO/human_GO.gpickle')
ontology_graph_literate = nx.relabel_nodes(ontology_graph, {key: data['name'] for key, data in ontology_graph.nodes(data=True)})
GO_columns = list(ontology_graph.nodes(data=True)['GO:0000001'].keys())
human_GO = pd.read_csv('/Users/alex/Documents/goenrich-web/GO/human_GO.csv')
human_GO['NCBI_ID'] = human_GO.NCBI_ID.str.split(';')
human_GO['ENSG'] = human_GO.ENSG.str.split(';')
def drop_nan_index_levels(df):
for i in list(range(df.index.nlevels))[::-1]:
if pd.isnull(df.index.get_level_values(i)).all():
df.index = df.index.droplevel(i)
return df
def KEGG_GO_venn(kegg_brite, GO_nodes):
KEGG_list = np.unique(kegg_brite.ENSG.dropna().values)
GO_list = np.unique(flatten(GO_nodes.ENSG.values))
return venn(KEGG_list, GO_list)
KEGG
proteasome_pathway_id = 'hsa03050'
proteasome_brite_id = 'ko03051'
Image(REST.kegg_get(proteasome_pathway_id, 'image').read())
proteasome_pathway_genes = get_KEGG_pathway_gene_members(proteasome_pathway_id)
proteasome_brite_genes = get_KEGG_BRITE_entry(proteasome_brite_id)
proteasome_brite_genes = proteasome_brite_genes.loc['Eukaryotic proteasome']
proteasome_brite_genes['in_KEGG_pathway'] = proteasome_brite_genes.ENSG.isin(proteasome_pathway_genes.ENSG).astype(int)
proteasome_brite_genes.to_pickle('targets/proteasome_brite.pickle')
GO
proteasome_subgraph, proteasome_nodes = GO_subgraph_from_query('proteasom')
draw_graph(proteasome_subgraph)
df = create_GO_by_gene_matrix(proteasome_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
proteasome_nodes.to_pickle('targets/proteasome_GO.pickle')
Overlap
KEGG_GO_venn(proteasome_brite_genes, proteasome_nodes)
KEGG
chaperones_brite_id = 'ko03110'
chaperones_brite_genes = get_KEGG_BRITE_entry(chaperones_brite_id)
chaperones_brite_genes.to_pickle('targets/chaperone_brite.pickle')
GO
chaperone_subgraph, chaperone_nodes = GO_subgraph_from_query('chaperone')
draw_graph(chaperone_subgraph)
df = create_GO_by_gene_matrix(chaperone_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
chaperone_nodes.to_pickle('targets/chaperone_GO.pickle')
Overlap
KEGG_GO_venn(chaperones_brite_genes, chaperone_nodes)
KEGG
autophagy_KEGG_pathway = 'hsa04140'
autophagy_KEGG_BRITE = 'ko04131'
Image(REST.kegg_get(autophagy_KEGG_pathway, 'image').read())
autophagy_pathway_genes = get_KEGG_pathway_gene_members(autophagy_KEGG_pathway)
autophagy_brite_genes = get_KEGG_BRITE_entry(autophagy_KEGG_BRITE)
autophagy_brite_genes = autophagy_brite_genes.loc['Autophagy']
autophagy_brite_genes['in_KEGG_pathway'] = autophagy_brite_genes.ENSG.isin(autophagy_pathway_genes.ENSG).astype(int)
autophagy_brite_genes.to_pickle('targets/autophagy_brite.pickle')
GO
autophagy_subgraph, autophagy_nodes = GO_subgraph_from_query('autophag')
df = create_GO_by_gene_matrix(autophagy_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
clustermap = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
autophagy_nodes.to_pickle('targets/autophagy_GO.pickle')
Overlap
KEGG_GO_venn(autophagy_brite_genes, autophagy_nodes)
KEGG
mitophagy_KEGG_pathway = 'hsa04137'
Image(REST.kegg_get(mitophagy_KEGG_pathway, 'image').read())
mitophagy_pathway_genes = get_KEGG_pathway_gene_members(mitophagy_KEGG_pathway)
mitophagy_brite_genes = pd.DataFrame(autophagy_brite_genes.loc['Mitophagy'])
mitophagy_brite_genes.drop(['in_KEGG_pathway'], axis=1, inplace=True)
mitophagy_brite_genes['in_KEGG_pathway'] = mitophagy_brite_genes.ENSG.isin(mitophagy_pathway_genes.ENSG).astype(int)
mitophagy_brite_genes.to_pickle('targets/mitophagy_brite.pickle')
GO
mitophagy_subgraph, mitophagy_nodes = GO_subgraph_from_query('mitophagy')
df = create_GO_by_gene_matrix(mitophagy_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
mitophagy_nodes.to_pickle('targets/mitophagy_GO.pickle')
Overlap
KEGG_GO_venn(mitophagy_brite_genes, mitophagy_nodes)
KEGG
lysosome_KEGG_pathway = 'hsa04142'
Image(REST.kegg_get(lysosome_KEGG_pathway, 'image').read())
lysosome_pathway_genes = get_KEGG_pathway_gene_members(lysosome_KEGG_pathway)
lysosome_pathway_genes.set_index('ENSG')['gene_name'].to_pickle('./targets/lysosome_kegg.pickle')
GO
lysosome_subgraph, lysosome_nodes = GO_subgraph_from_query('lysosom')
df = create_GO_by_gene_matrix(lysosome_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
lysosome_nodes.to_pickle('targets/lysosome_GO.pickle')
Overlap
KEGG_GO_venn(lysosome_pathway_genes, lysosome_nodes)
brite = 'ko03036'
brite_genes = get_KEGG_BRITE_entry(brite)
brite_genes = brite_genes.loc['Eukaryotic type']
# Silencing chromatin modifiers,
# TFs which recognize ERVs
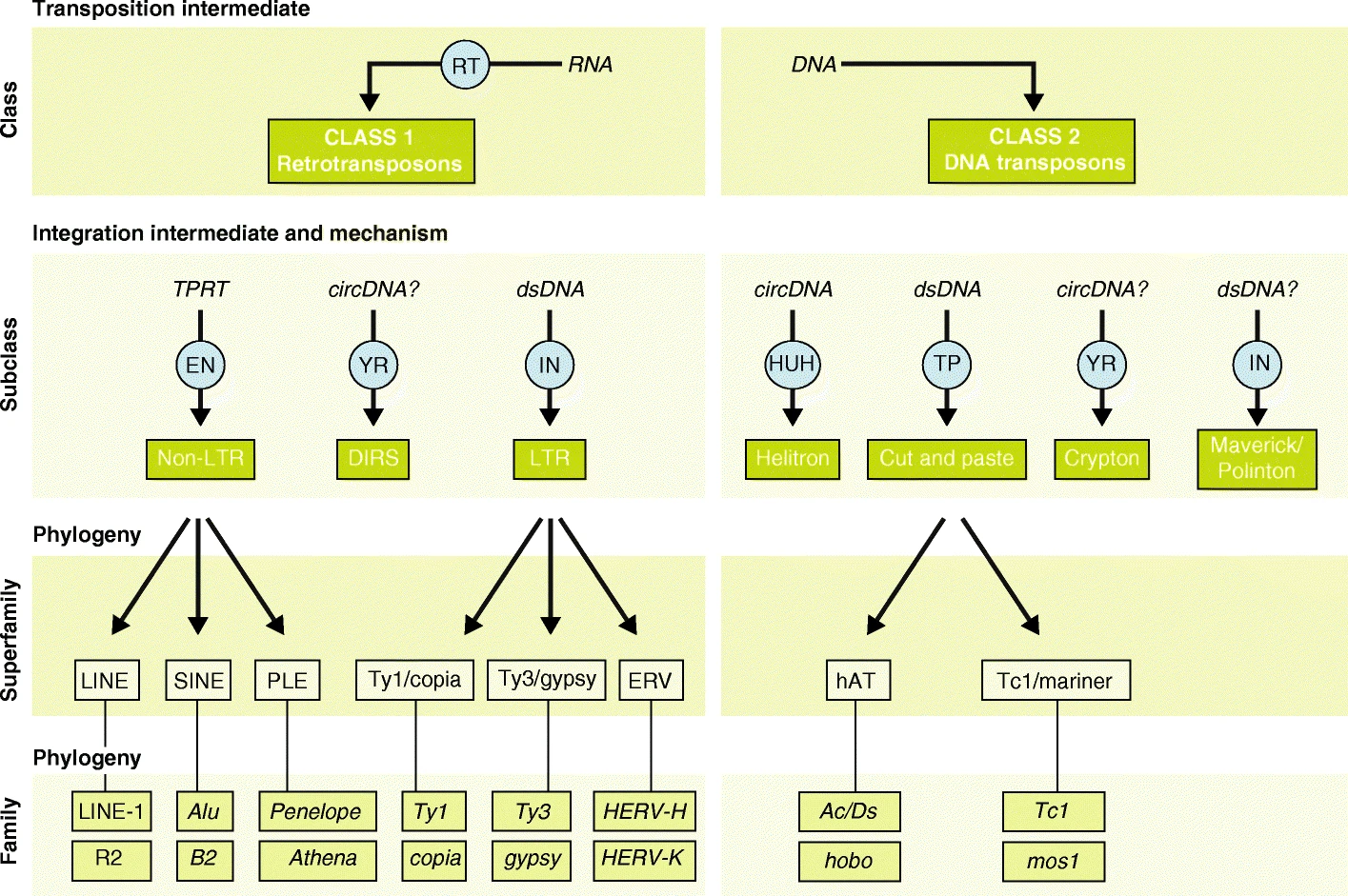
KEGG
gene_silencing_brite = brite_genes.loc['Gene silencing']
gene_silencing_brite.index = gene_silencing_brite.index.droplevel(2)
gene_silencing_brite.to_pickle('targets/gene_silencing_brite.pickle')
GO
transposon_subgraph, transposon_nodes = GO_subgraph_from_query('transpos')
draw_graph(transposon_subgraph)
df = create_GO_by_gene_matrix(transposon_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
transposon_nodes.to_pickle('targets/transposon_GO.pickle')
Overlap
KEGG_GO_venn(gene_silencing_brite, transposon_nodes)
KEGG
chromatin_remodeling_brite = brite_genes.loc['Chromatin remodeling factors']
chromatin_remodeling_brite.index = chromatin_remodeling_brite.index.droplevel(2).droplevel(1)
chromatin_remodeling_brite.to_pickle('targets/chromatin_remodeling_brite.pickle')
histone_modification_brite = brite_genes.loc['Histone modification proteins']
histone_modification_brite = drop_nan_index_levels(histone_modification_brite)
histone_modification_brite.to_pickle('targets/histone_modification_brite.pickle')
GO
chromatin_remodeling_subgraph, chromatin_remodeling_nodes = GO_subgraph_from_query('chromatin remodel')
draw_graph(chromatin_remodeling_subgraph)
chromatin_remodeling_nodes.to_pickle('targets/chromatin_remodeling_GO.pickle')
Overlap
KEGG_GO_venn(chromatin_remodeling_brite, chromatin_remodeling_nodes)
KEGG
heterochromatin_formation_brite = brite_genes.loc['Heterochromatin formation proteins']
heterochromatin_formation_brite = drop_nan_index_levels(heterochromatin_formation_brite)
heterochromatin_formation_brite.to_pickle('targets/heterochromatin_formation_brite.pickle')
GO
heterochromatin_subgraph, heterochromatin_nodes = GO_subgraph_from_query('heterochromatin')
draw_graph(heterochromatin_subgraph)
heterochromatin_nodes.to_pickle('targets/heterochromatin_GO.pickle')
Overlap
KEGG_GO_venn(heterochromatin_formation_brite, heterochromatin_nodes)
KEGG
nucleosome_assembly_brite = brite_genes.loc['Nucleosome assembly factors']
nucleosome_assembly_brite = drop_nan_index_levels(nucleosome_assembly_brite)
nucleosome_assembly_brite.to_pickle('targets/nucleosome_assembly_brite.pickle')
GO
nucleosome_assembly_subgraph, nucleosome_assembly_nodes = GO_subgraph_from_query('nucleosome assembly')
draw_graph(nucleosome_assembly_subgraph)
nucleosome_assembly_nodes.to_pickle('targets/nucleosome_assembly_GO.pickle')
Overlap
KEGG_GO_venn(nucleosome_assembly_brite, nucleosome_assembly_nodes)
Image('https://upload.wikimedia.org/wikipedia/commons/b/b4/Structure_and_function_of_the_nuclear_lamina.jpg')
KEGG
nuclear_lamin_brite_id = 'ko04812+K12641'
nuclear_lamin_brite = get_KEGG_BRITE_entry(nuclear_lamin_brite_id)
nuclear_lamin_brite = nuclear_lamin_brite.loc['Eukaryotic cytoskeleton proteins', 'Intermediate filaments', 'Intermediate filaments', 'Type V: Nuclear lamins']
nuclear_lamin_brite.index = ['Nuclear lamins']*len(nuclear_lamin_brite)
nuclear_lamin_brite.to_pickle('targets/nuclear_lamin_brite.pickle')
GO
nuclear_lamin_subgraph, nuclear_lamin_nodes = GO_subgraph_from_query('nuclear lamin')
draw_graph(nuclear_lamin_subgraph)
nuclear_lamin_nodes.to_pickle('targets/nuclear_lamin_GO.pickle')
Overlap
KEGG_GO_venn(nuclear_lamin_brite, nuclear_lamin_nodes)
KEGG
cell_cycle_pathway_id = 'hsa04110'
Image(REST.kegg_get(cell_cycle_pathway_id, 'image').read())
cell_cycle_pathway_genes = get_KEGG_pathway_gene_members(cell_cycle_pathway_id)
cell_cycle_pathway_genes.to_pickle('cell_cycle_pathway.pickle')
GO
cell_cycle_subgraph, cell_cycle_nodes = GO_subgraph_from_query('cell cycle')
df = create_GO_by_gene_matrix(cell_cycle_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
matplotlib_inline.backend_inline.set_matplotlib_formats('png')
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
draw_graph(cell_cycle_subgraph)
cell_cycle_nodes.to_pickle('targets/cell_cycle_GO.pickle')
Overlap
KEGG_GO_venn(cell_cycle_pathway_genes, cell_cycle_nodes)
pd.set_option('display.max_rows', 200)
matplotlib_inline.backend_inline.set_matplotlib_formats('svg')
KEGG
chemokine_pathway_id = 'hsa04062'
TNF_pathway_id = 'hsa04668'
complement_pathway_id = 'hsa04610'
leukocyte_transendothelial_migration_pathway_id = 'hsa04670'
Image(REST.kegg_get(chemokine_pathway_id, 'image').read())
Image(REST.kegg_get(TNF_pathway_id, 'image').read())
Image(REST.kegg_get(complement_pathway_id, 'image').read())
Image(REST.kegg_get(leukocyte_transendothelial_migration_pathway_id, 'image').read())
chemokine_pathway_genes = get_KEGG_pathway_gene_members(chemokine_pathway_id)
TNF_pathway_genes = get_KEGG_pathway_gene_members(TNF_pathway_id)
complement_pathway_genes = get_KEGG_pathway_gene_members(complement_pathway_id)
leukocyte_transendothelial_migration_pathway_genes = get_KEGG_pathway_gene_members(leukocyte_transendothelial_migration_pathway_id)
chemokine_pathway_genes.index = ['Chemokine signaling pathway']*len(chemokine_pathway_genes)
TNF_pathway_genes.index = ['TNF signaling pathway']*len(TNF_pathway_genes)
complement_pathway_genes.index = ['Complement and coagulation cascades']*len(complement_pathway_genes)
leukocyte_transendothelial_migration_pathway_genes.index = ['Leukocyte transendothelial migration']*len(leukocyte_transendothelial_migration_pathway_genes)
inflammation_genes = pd.concat((chemokine_pathway_genes,TNF_pathway_genes,complement_pathway_genes,leukocyte_transendothelial_migration_pathway_genes))
inflammation_genes.to_pickle('targets/inflammation_brite.pickle')
GO
inflammation_subgraph, inflammation_nodes = GO_subgraph_from_query('inflammat')
df = create_GO_by_gene_matrix(inflammation_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
matplotlib_inline.backend_inline.set_matplotlib_formats('png')
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
draw_graph(inflammation_subgraph)
inflammation_nodes.to_pickle('targets/inflammation_GO.pickle')
Overlap
matplotlib_inline.backend_inline.set_matplotlib_formats('svg')
KEGG_GO_venn(inflammation_genes, inflammation_nodes)
KEGG
cytokines_KEGG_BRITE = 'ko04052'
cytokines_brite_genes = get_KEGG_BRITE_entry(cytokines_KEGG_BRITE)
cytokines_brite_genes.to_pickle('targets/cytokines_brite.pickle')
GO
cytokines_subgraph, cytokines_nodes = GO_subgraph_from_query('cytokine')
df = create_GO_by_gene_matrix(cytokines_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
matplotlib_inline.backend_inline.set_matplotlib_formats('png')
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
draw_graph(cytokines_subgraph)
cytokines_nodes.to_pickle('targets/cytokines_GO.pickle')
Overlap
matplotlib_inline.backend_inline.set_matplotlib_formats('svg')
KEGG_GO_venn(cytokines_brite_genes, cytokines_nodes)
KEGG
cGAS_STING_pathway_id = 'hsa04623'
Image(REST.kegg_get(cGAS_STING_pathway_id, 'image').read())
cGAS_STING_pathway_genes = get_KEGG_pathway_gene_members(cGAS_STING_pathway_id)
cGAS_STING_pathway_genes.to_pickle('targets/cGAS_STING_kegg.pickle')
GO
cGAS_STING_subgraph, cGAS_STING_nodes = GO_subgraph_from_query('activation of innate immune response')
draw_graph(cGAS_STING_subgraph)
cGAS_STING_nodes.to_pickle('targets/cGAS_STING_GO.pickle')
Overlap
KEGG_GO_venn(cGAS_STING_pathway_genes, cGAS_STING_nodes)
KEGG
senescence_pathway = 'hsa04218'
Image(REST.kegg_get(senescence_pathway, 'image').read())
senescence_pathway_genes = get_KEGG_pathway_gene_members(senescence_pathway)
senescence_pathway_genes.to_pickle('targets/senescence_kegg.pickle')
GO
senescence_subgraph, senescence_nodes = GO_subgraph_from_query('senescence')
df = create_GO_by_gene_matrix(senescence_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
matplotlib_inline.backend_inline.set_matplotlib_formats('svg')
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
draw_graph(senescence_subgraph)
senescence_nodes.to_pickle('targets/senescence_GO.pickle')
Overlap
KEGG_GO_venn(senescence_pathway_genes, senescence_nodes)
KEGG
inflammasome_pathway = 'hsa04621'
Image(REST.kegg_get(inflammasome_pathway, 'image').read())
inflammasome_pathway_genes = get_KEGG_pathway_gene_members(inflammasome_pathway)
inflammasome_pathway_genes.to_pickle('targets/inflammasome_kegg.pickle')
GO
inflammasome_subgraph, inflammasome_nodes = GO_subgraph_from_query('inflammasome')
df = create_GO_by_gene_matrix(inflammasome_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
# matplotlib_inline.backend_inline.set_matplotlib_formats('svg')
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
draw_graph(inflammasome_subgraph)
inflammasome_nodes.to_pickle('targets/inflammasome_GO.pickle')
Overlap
KEGG_GO_venn(inflammasome_pathway_genes, inflammasome_nodes)
KEGG
glycolysis_pathway_id = 'hsa00010'
# glycolysis_brite_id = 'ko03051'
Image(REST.kegg_get(glycolysis_pathway_id, 'image').read())
glycolysis_pathway_genes = get_KEGG_pathway_gene_members(glycolysis_pathway_id)
# proteasome_brite_genes = proteasome_brite_genes.loc['Eukaryotic proteasome']
# proteasome_brite_genes['in_KEGG_pathway'] = proteasome_brite_genes.ENSG.isin(proteasome_pathway_genes.ENSG).astype(int)
glycolysis_pathway_genes.to_pickle('targets/glycolysis_kegg.pickle')
GO
glycolysis_subgraph, glycolysis_nodes = GO_subgraph_from_query('glycolysis')
df = create_GO_by_gene_matrix(glycolysis_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
draw_graph(glycolysis_subgraph)
glycolysis_nodes.to_pickle('targets/glycolysis_GO.pickle')
Overlap
KEGG_GO_venn(glycolysis_pathway_genes, glycolysis_nodes)
KEGG
tca_pathway_id = 'hsa00020'
# tca_brite_id = 'ko03051'
Image(REST.kegg_get(tca_pathway_id, 'image').read())
tca_pathway_genes = get_KEGG_pathway_gene_members(tca_pathway_id)
# proteasome_brite_genes = proteasome_brite_genes.loc['Eukaryotic proteasome']
# proteasome_brite_genes['in_KEGG_pathway'] = proteasome_brite_genes.ENSG.isin(proteasome_pathway_genes.ENSG).astype(int)
tca_pathway_genes.to_pickle('targets/tca_kegg.pickle')
GO
tca_subgraph, tca_nodes = GO_subgraph_from_query('tricarboxylic acid cycle')
df = create_GO_by_gene_matrix(tca_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
draw_graph(tca_subgraph)
tca_nodes.to_pickle('targets/tca_GO.pickle')
Overlap
KEGG_GO_venn(tca_pathway_genes, tca_nodes)
KEGG
ETC_pathway = 'hsa00190'
Image(REST.kegg_get(ETC_pathway, 'image').read())
ETC_pathway_genes = get_KEGG_pathway_gene_members(ETC_pathway)
ETC_pathway_genes.to_pickle('targets/ETC_kegg.pickle')
GO
ETC_subgraph, ETC_nodes = GO_subgraph_from_query('electron transport chain')
df = create_GO_by_gene_matrix(ETC_nodes, attr='gene_name')
df = sort_df_by_hclust_olo(df)
ax = plot_gene_membership_in_GO_terms(df, figsize=(6, 10))
draw_graph(ETC_subgraph)
ETC_nodes.to_pickle('targets/ETC_GO.pickle')
Overlap
KEGG_GO_venn(ETC_pathway_genes, ETC_nodes)
KEGG
pd.set_option('display.max_rows', 200)
mitochondrial_biogenesis_brite_id = 'ko03029'
mitochondrial_biogenesis_brite_genes = get_KEGG_BRITE_entry(mitochondrial_biogenesis_brite_id)
mitochondrial_biogenesis_brite_genes = mitochondrial_biogenesis_brite_genes.loc[['Mitochondrial DNA transcription, translation, and replication factors','Mitochondrial quality control factors']]
# mitochondrial_biogenesis_brite.loc['Mitochondrial DNA transcription, translation, and replication factors']
mitochondrial_biogenesis_brite_genes.to_pickle('targets/mitochondrial_biogenesis_brite.pickle')
GO
mitochondrial_biogenesis_subgraph, mitochondrial_biogenesis_nodes = GO_subgraph_from_query('mitochondrial fission')
draw_graph(mitochondrial_biogenesis_subgraph)
mitochondrial_biogenesis_nodes.to_pickle('targets/mitochondrial_biogenesis_GO.pickle')
Overlap
KEGG_GO_venn(mitochondrial_biogenesis_brite_genes, mitochondrial_biogenesis_nodes)
GO
oxidative_stress_subgraph, oxidative_stress_nodes = GO_subgraph_from_query('oxidative stress')
draw_graph(oxidative_stress_subgraph)
oxidative_stress_nodes.to_pickle('targets/oxidative_stress_GO.pickle')